Plotting#
%load_ext autoreload
%autoreload 2
import logging
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import anndata as ad
from alphapepttools import pp
from alphapepttools.pl import colors
from alphapepttools.pl.colors import show_rgba_color_list
from alphapepttools.pl.figure import create_figure, label_axes, save_figure
from alphapepttools.pl.plots import Plots
logging.basicConfig(level=logging.INFO)
Basic colors, palettes and color maps for alphapepttools#
# colors, derived from the basic palette
base_colors = [
"red",
"green",
"blue",
"orange",
"yellow",
"lightred",
"lightgreen",
"lightblue",
"lightorange",
"grey",
"white",
"black",
]
show_rgba_color_list([colors.BaseColors.get(color) for color in base_colors])
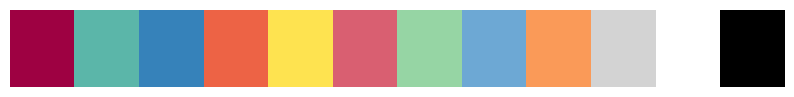
# color palettes
palettes = ["binary", "qualitative"]
for palette in palettes:
show_rgba_color_list(colors.BasePalettes.get(palette))


# colormaps
maps = ["sequential", "diverging"]
for cmap in maps:
show_rgba_color_list(list(colors.BaseColormaps.get(cmap)(np.arange(0, 1, 0.001))))


test_data = [*list(np.random.rand(100)), -100, 100]
# map colormaps to numerical values without capping
mapped_colors = colors.MappedColormaps("sequential").fit_transform(test_data)
show_rgba_color_list(mapped_colors)
# map colormaps to numerical values with capping
mapped_colors = colors.MappedColormaps("sequential", (5, 95)).fit_transform(test_data)
show_rgba_color_list(mapped_colors)


Demonstrate figure.py submodule style & label handling#
# Create a 2x3 grid of subplots in an AxisManager
fig, axm = create_figure(
nrows=2,
ncols=3,
figsize=(10, 6),
# figure_padding=3
)
# Example dataset
x = np.linspace(0, 10, 100)
y_funcs = [
lambda x: np.sin(x),
lambda x: np.cos(x),
lambda x: np.tan(x),
lambda x: x**2,
lambda x: np.exp(x / 5),
lambda x: np.log(x + 1),
]
# Get qualitative palette
palette = colors.BasePalettes.get("qualitative", len(y_funcs))
# Iterate through all axes using next() and plot different functions
try:
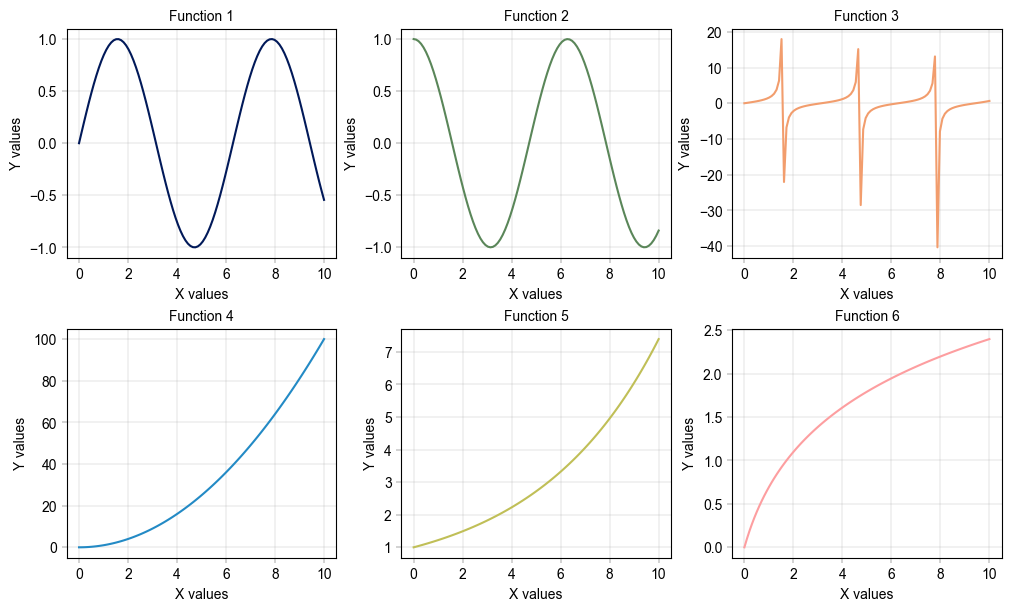
for i, func in enumerate(y_funcs):
ax = axm.next()
ax.plot(x, func(x), color=palette[i])
label_axes(ax, xlabel="X values", ylabel="Y values", title=f"Function {i + 1}")
except StopIteration:
pass
plt.show()
# Save the figure
save_figure(
fig=fig,
filename="example_figure.png",
output_dir="./example_outputs",
dpi=300,
transparent=False,
)
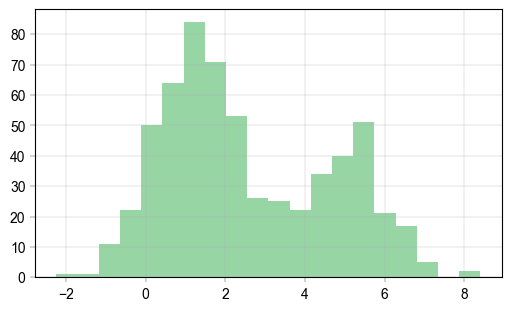
Basic histogram#
example_df = pd.DataFrame(
{
"values": np.concatenate([np.random.normal(i, size=200) + np.random.normal(i) for i in range(3)]),
"values2": np.concatenate([np.random.normal(i, size=200) + np.random.normal(i) for i in range(3)]),
"levels": [i for i in range(3) for _ in range(200)],
"levels2": np.arange(0, 600),
}
)
# with one color
fig, axm = create_figure(figsize=(5, 3))
# Use the AxisManager
ax = axm.next()
Plots.histogram(
data=example_df,
value_column="values",
bins=20,
color="lightgreen",
ax=ax,
)
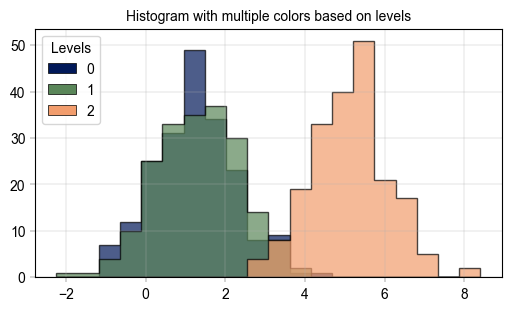
# with multiple colors based on levels
fig, axm = create_figure(figsize=(5, 3))
# Apply the AxisManager
ax = axm.next()
palette = colors.BasePalettes.get("qualitative", example_df["levels"].nunique())
Plots.histogram(
data=example_df,
value_column="values",
color_map_column="levels",
palette=palette,
bins=20,
color="blue",
ax=ax,
legend="auto",
hist_kwargs={"alpha": 0.7, "histtype": "stepfilled", "edgecolor": "k"},
legend_kwargs={"title": "Levels", "loc": "upper left"},
)
label_axes(ax, title="Histogram with multiple colors based on levels")
plt.show()
# Save the figure
save_figure(
fig=fig,
filename="example_histogram.png",
output_dir="./example_outputs",
dpi=300,
transparent=False,
)
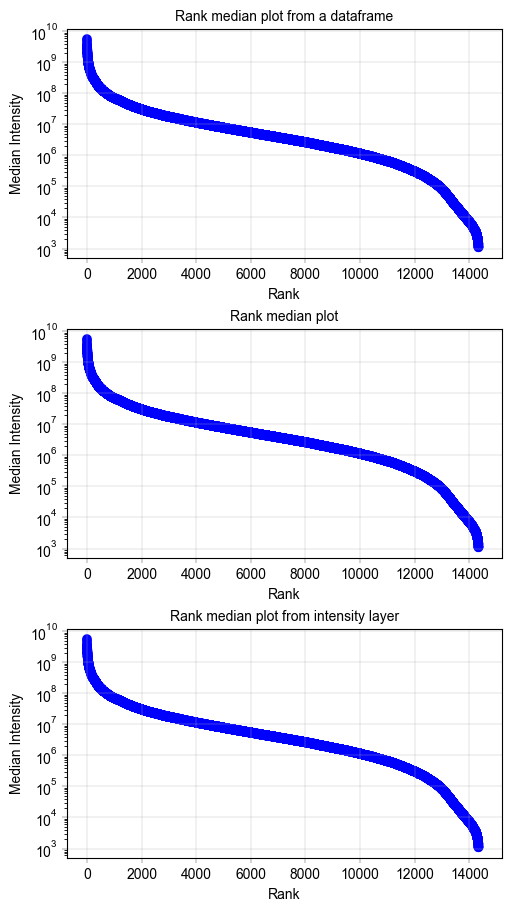
Rank plot of protein median intensities across all samples#
def load_diann_pg_matrix(
data_path: str,
) -> ad.AnnData:
"""Load diann sample data into a pandas dataframe"""
from alphapepttools.pp.data import _to_anndata
X = pd.read_pickle(data_path)
# to be replaced by AlphaBase PSM reader
return _to_anndata(X)
# read an example proteomics data
# load protein groups into an AnnData object with index & columns as obs & var
data_path = "./example_data/HeLa_QC_data.pkl"
adata = load_diann_pg_matrix(data_path)
# Add sample metadata
sample_metadata = pd.read_csv("./example_data/HeLa_QC_sample_metadata.csv", index_col=0)
adata = pp.add_metadata(adata, sample_metadata, axis=0)
# Add feature metadata
feature_metadata = pd.read_csv("./example_data/HeLa_QC_feature_metadata.csv", index_col=0)
adata = pp.add_metadata(adata, feature_metadata, axis=1)
Add a layer called ‘intensity’ to store raw data
adata.layers["intensity"] = adata.X.copy()
fig, axm = create_figure(3, 1, figsize=(5, 9))
# Create median rank plot from a dataframe
ax = axm.next()
data_df = adata.to_df()
Plots.rank_median_plot(data_df, ax=ax)
label_axes(ax, title="Rank median plot from a dataframe")
# figure using the default settings for AnnData
ax = axm.next()
Plots.rank_median_plot(adata, ax=ax)
label_axes(ax, title="Rank median plot")
# figure with a non-default layer
ax = axm.next()
Plots.rank_median_plot(adata, layer="intensity", ax=ax)
label_axes(ax, title="Rank median plot from intensity layer")
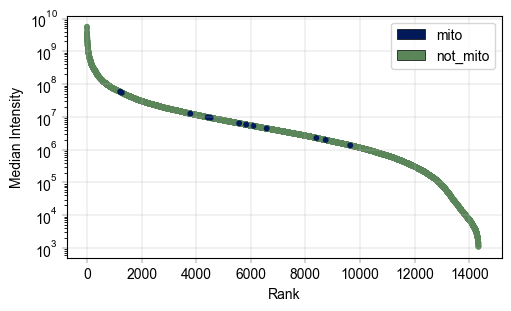
# create a column for mitochondrial proteins tp color code the rank plot
mt_mask = adata.var["Genes"].str.startswith("MT-")
adata.var["Mito_prot"] = "not_mito"
adata.var.loc[mt_mask, "Mito_prot"] = "mito"
fig, axs = create_figure(figsize=(5, 3))
ax = axs.next()
Plots.rank_median_plot(adata, ax=ax, color_map_column="Mito_prot", scatter_kwargs={"s": 10}, legend="auto")
# get median intensity of all proteins and color by above and below median of medians
medians = np.nanmedian(adata.X, axis=0)
median_median = np.nanmedian(medians)
adata.var["Median_intensity"] = medians
med_mask = adata.var["Median_intensity"] > median_median
adata.var["Above_median"] = "blue"
adata.var.loc[med_mask, "Above_median"] = "red"
fig, axs = create_figure(figsize=(5, 3))
ax = axs.next()
Plots.rank_median_plot(adata, ax=ax, color_map_column="Above_median", scatter_kwargs={"s": 10})
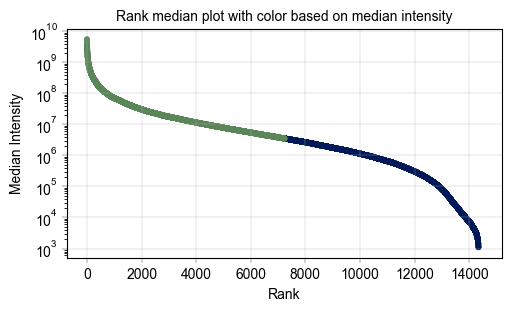
label_axes(ax, title="Rank median plot with color based on median intensity")