Cellpose Segmentation on human protein atlas images#
[2]:
import os
from scportrait.pipeline.featurization import CellFeaturizer
from scportrait.pipeline.extraction import HDF5CellExtraction
from scportrait.pipeline.project import Project
from scportrait.pipeline.segmentation.workflows import CytosolSegmentationCellpose
from scportrait.pipeline.selection import LMDSelection
import scportrait
[3]:
project_location = "project"
config_path = scportrait.data.get_config_file(config_id = "dataset_2_config")
project = Project(
os.path.abspath(project_location),
config_path=config_path,
overwrite=True,
debug=True,
segmentation_f=CytosolSegmentationCellpose,
extraction_f=HDF5CellExtraction,
featurization_f=CellFeaturizer,
selection_f=LMDSelection,
)
Downloading...: 1.57kB [00:00, 2.48MB/s]
Updating project config file.
[10/04/2025 19:03:09] Loading config from /Users/sophia/Documents/GitHub/scPortrait/examples/notebooks/example_projects/example_2/project/config.yml
[10/04/2025 19:03:09] Compression algorithm for extracted single-cell images: lzf
[10/04/2025 19:03:09] No cache directory specified in config using current working directory /Users/sophia/Documents/GitHub/scPortrait/examples/notebooks/example_projects/example_2.
[10/04/2025 19:03:09] No cache directory specified in config using current working directory /Users/sophia/Documents/GitHub/scPortrait/examples/notebooks/example_projects/example_2.
[4]:
dataset_2_path = scportrait.data.dataset_2()
# these example images are downloaded from the human protein atlas (www.proteinatlas.org)
images = [f"{dataset_2_path}/Ch1.tif", f"{dataset_2_path}/Ch2.tif", f"{dataset_2_path }/Ch3.tif"]
project.load_input_from_tif_files(images, channel_names = ["Channel1", "Channel2", "Channel3"])
[10/04/2025 19:03:09] Output location /Users/sophia/Documents/GitHub/scPortrait/examples/notebooks/example_projects/example_2/project/scportrait.sdata already exists. Overwriting.
INFO The Zarr backing store has been changed from None the new file path:
/Users/sophia/Documents/GitHub/scPortrait/examples/notebooks/example_projects/example_2/project/scportrait
.sdata
[10/04/2025 19:03:09] Initialized temporary directory at /Users/sophia/Documents/GitHub/scPortrait/examples/notebooks/example_projects/example_2/Project_hdt2zmjo for Project
[10/04/2025 19:03:09] Image input_image written to sdata object.
[10/04/2025 19:03:10] Cleaned up temporary directory at <TemporaryDirectory '/Users/sophia/Documents/GitHub/scPortrait/examples/notebooks/example_projects/example_2/Project_hdt2zmjo'>
[5]:
project.plot_input_image()
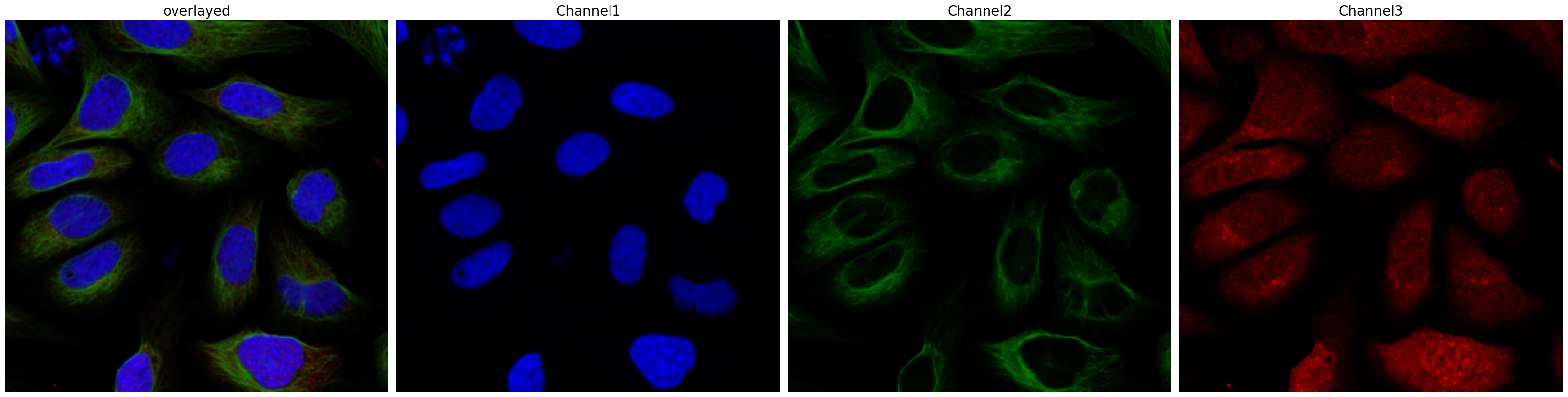
[6]:
project.segment()
[10/04/2025 19:03:13] GPU Status for segmentation is True and will segment using the following device mps.
[10/04/2025 19:03:13] Segmenting nucleus using the following model: nuclei
[10/04/2025 19:03:16] Segmenting cytosol using the following model: cyto2
[10/04/2025 19:03:16] Performing filtering to match Cytosol and Nucleus IDs.
[10/04/2025 19:03:16] Removed 4 nuclei and 2 cytosols due to filtering.
[10/04/2025 19:03:16] After filtering, 7 matching nuclei and cytosol masks remain.
[10/04/2025 19:03:17] Total time to perform nucleus and cytosol mask matching filtering: 0.59 seconds
[10/04/2025 19:03:17] Segmentation seg_all_nucleus written to sdata object.
[10/04/2025 19:03:18] Points centers_seg_all_nucleus written to sdata object.
[10/04/2025 19:03:18] Segmentation seg_all_cytosol written to sdata object.
[10/04/2025 19:03:19] Points centers_seg_all_cytosol written to sdata object.
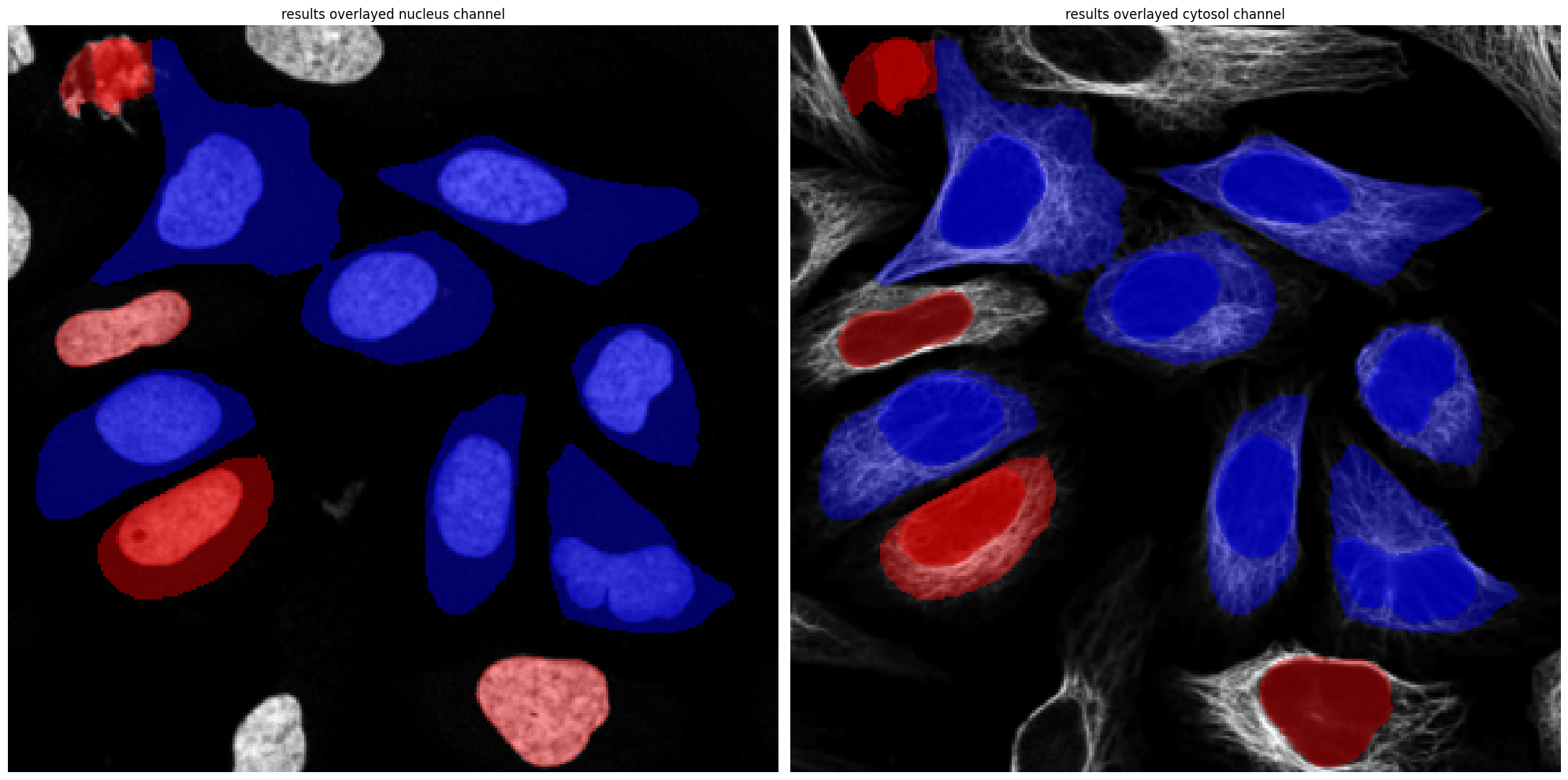
[7]:
project.plot_segmentation_masks()
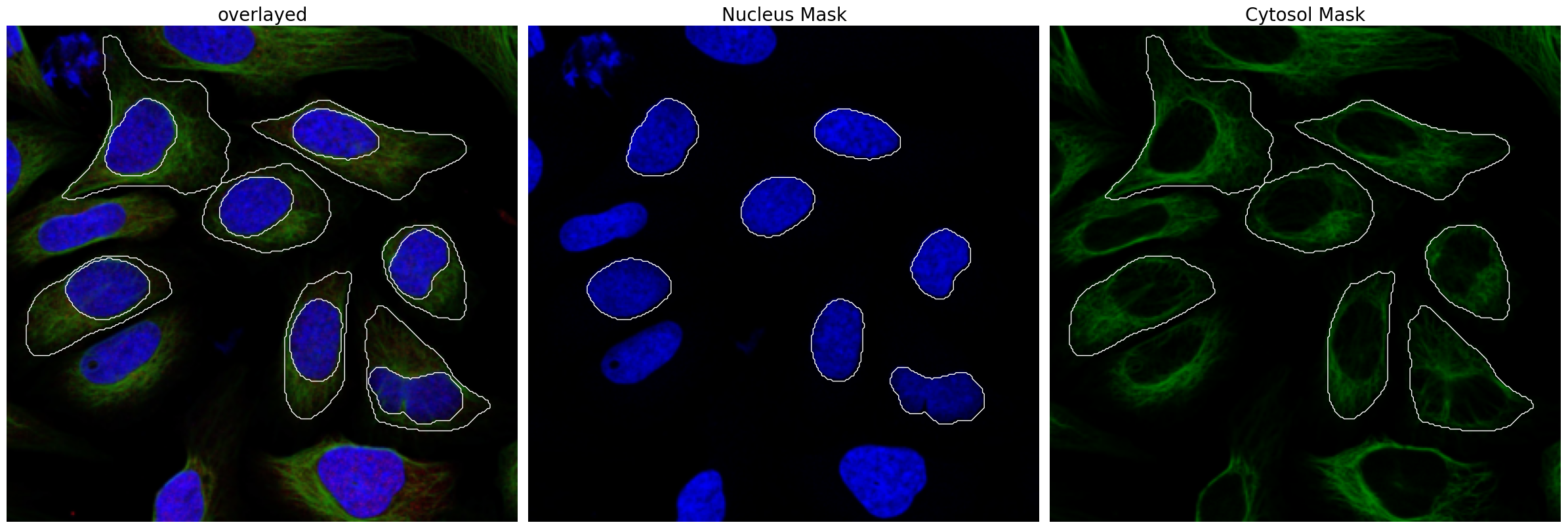
[8]:
project.extract()
[10/04/2025 19:03:20] Initialized temporary directory at /var/folders/35/p4c58_4n3bb0bxnzgns1t7kh0000gn/T/./HDF5CellExtraction_h_kn4vdv for HDF5CellExtraction
[10/04/2025 19:03:20] Created new directory for extraction results: /Users/sophia/Documents/GitHub/scPortrait/examples/notebooks/example_projects/example_2/project/extraction/data
[10/04/2025 19:03:20] Setup output folder at /Users/sophia/Documents/GitHub/scPortrait/examples/notebooks/example_projects/example_2/project/extraction/data
[10/04/2025 19:03:20] Found 2 segmentation masks for the given key in the sdata object. Will be extracting single-cell images based on these masks: ['seg_all_nucleus', 'seg_all_cytosol']
[10/04/2025 19:03:20] Using seg_all_nucleus as the main segmentation mask to determine cell centers.
[10/04/2025 19:03:20] A total of 0 cells were too close to the image border to be extracted. Their cell_ids were saved to file /Users/sophia/Documents/GitHub/scPortrait/examples/notebooks/example_projects/example_2/project/extraction/removed_classes.csv.
[10/04/2025 19:03:20] Container for single-cell data created.
[10/04/2025 19:03:20] Extraction Details:
[10/04/2025 19:03:20] --------------------------------
[10/04/2025 19:03:20] Number of input image channels: 3
[10/04/2025 19:03:20] Number of segmentation masks used during extraction: 2
[10/04/2025 19:03:20] Number of generated output images per cell: 5
[10/04/2025 19:03:20] Number of unique cells to extract: 7
[10/04/2025 19:03:20] Extracted Image Dimensions: 110 x 110
[10/04/2025 19:03:20] Normalization of extracted images: True
[10/04/2025 19:03:20] Percentile normalization range for single-cell images: [0.01, 0.99]
[10/04/2025 19:03:20] Starting single-cell image extraction of 7 cells...
[10/04/2025 19:03:20] Loading input images to memory mapped arrays...
[10/04/2025 19:03:21] Finished transferring data to memory mapped arrays. Time taken: 0.19 seconds.
[10/04/2025 19:03:21] Using batch size of 100 for multiprocessing.
[10/04/2025 19:03:21] Running in multiprocessing mode with 1 threads.
[10/04/2025 19:03:21] Finished extraction in 0.15 seconds (47.90 cells / second)
[10/04/2025 19:03:21] Benchmarking times saved to file.
[10/04/2025 19:03:21] Cleaned up temporary directory at <TemporaryDirectory '/var/folders/35/p4c58_4n3bb0bxnzgns1t7kh0000gn/T/./HDF5CellExtraction_h_kn4vdv'>
[9]:
project.plot_single_cell_images()
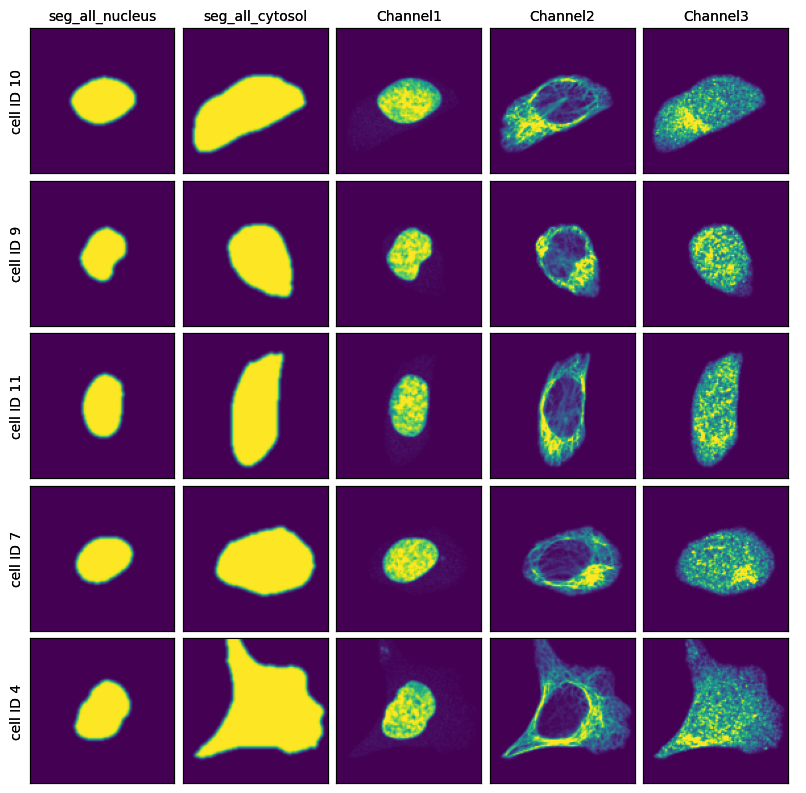
[10]:
project.featurize(overwrite=True)
Using extraction directory: /Users/sophia/Documents/GitHub/scPortrait/examples/notebooks/example_projects/example_2/project/extraction/data/single_cells.h5sc
[10/04/2025 19:03:22] Initialized temporary directory at /Users/sophia/Documents/GitHub/scPortrait/examples/notebooks/example_projects/example_2/CellFeaturizer_47gg1_ai for CellFeaturizer
[10/04/2025 19:03:22] Started CellFeaturization of all available channels.
[10/04/2025 19:03:22] Overwrite flag is set, deleting existing directory for featurization results.
[10/04/2025 19:03:22] Created new directory for featurization results: /Users/sophia/Documents/GitHub/scPortrait/examples/notebooks/example_projects/example_2/project/featurization/complete_CellFeaturizer
[10/04/2025 19:03:22] CPU specified in config file but MPS available on system. Consider changing the device for the next run.
[10/04/2025 19:03:22] Initialized temporary directory at /Users/sophia/Documents/GitHub/scPortrait/examples/notebooks/example_projects/example_2/CellFeaturizer_gfwyy78f for CellFeaturizer
[10/04/2025 19:03:22] Reading data from path: /Users/sophia/Documents/GitHub/scPortrait/examples/notebooks/example_projects/example_2/project/extraction/data/single_cells.h5sc
[10/04/2025 19:03:22] Processing dataset with 7 cells
[10/04/2025 19:03:22] Dataloader generated with a batchsize of 900 and 0 workers. Dataloader contains 1 entries.
[10/04/2025 19:03:22] Started processing of 1 batches.
[10/04/2025 19:03:22] finished processing.
[10/04/2025 19:03:22] Results saved to file: /Users/sophia/Documents/GitHub/scPortrait/examples/notebooks/example_projects/example_2/project/featurization/complete_CellFeaturizer/calculated_image_features.csv
[10/04/2025 19:03:22] Table CellFeaturizer_nucleus written to sdata object.
[10/04/2025 19:03:22] Table CellFeaturizer_cytosol written to sdata object.
[10/04/2025 19:03:22] GPU memory before performing cleanup: None
[10/04/2025 19:03:22] GPU memory after performing cleanup: None
[10/04/2025 19:03:22] Cleaned up temporary directory at <TemporaryDirectory '/Users/sophia/Documents/GitHub/scPortrait/examples/notebooks/example_projects/example_2/CellFeaturizer_gfwyy78f'>
[11]:
# load classification results
results = project.sdata['CellFeaturizer_cytosol'].to_df().merge(project.sdata['CellFeaturizer_cytosol'].obs, left_index=True, right_index=True).drop(columns = "region")
results
[11]:
| nucleus_area | cytosol_area | cytosol_only_area | Channel1_mean_nucleus | Channel1_median_nucleus | Channel1_quant75_nucleus | Channel1_quant25_nucleus | Channel1_summed_intensity_nucleus | Channel1_summed_intensity_area_normalized_nucleus | Channel1_mean_cytosol | ... | Channel3_quant25_cytosol | Channel3_summed_intensity_cytosol | Channel3_summed_intensity_area_normalized_cytosol | Channel3_mean_cytosol_only | Channel3_median_cytosol_only | Channel3_quant75_cytosol_only | Channel3_quant25_cytosol_only | Channel3_summed_intensity_cytosol_only | Channel3_summed_intensity_area_normalized_cytosol_only | scportrait_cell_id | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 2043.0 | 6255.0 | 4212.0 | 0.093840 | 0.0 | 0.024887 | 0.0 | 1135.460449 | 0.269578 | 0.093840 | ... | 0.0 | 2101.005859 | 0.498814 | 0.173637 | 0.0 | 0.315674 | 0.0 | 2101.005859 | 0.498814 | 4 |
| 1 | 1746.0 | 4319.0 | 2573.0 | 0.073899 | 0.0 | 0.007010 | 0.0 | 894.177795 | 0.347523 | 0.073899 | ... | 0.0 | 1735.827393 | 0.674632 | 0.143457 | 0.0 | 0.119019 | 0.0 | 1735.827393 | 0.674632 | 5 |
| 2 | 1768.0 | 3839.0 | 2071.0 | 0.079282 | 0.0 | 0.001667 | 0.0 | 959.311768 | 0.463212 | 0.079282 | ... | 0.0 | 1442.796143 | 0.696666 | 0.119239 | 0.0 | 0.028942 | 0.0 | 1442.796143 | 0.696666 | 7 |
| 3 | 1586.0 | 2841.0 | 1255.0 | 0.067117 | 0.0 | 0.000000 | 0.0 | 812.111816 | 0.647101 | 0.067117 | ... | 0.0 | 1124.324585 | 0.895876 | 0.092919 | 0.0 | 0.000000 | 0.0 | 1124.324585 | 0.895876 | 9 |
| 4 | 1972.0 | 4059.0 | 2087.0 | 0.086109 | 0.0 | 0.003860 | 0.0 | 1041.914307 | 0.499240 | 0.086109 | ... | 0.0 | 1476.587402 | 0.707517 | 0.122032 | 0.0 | 0.053192 | 0.0 | 1476.587402 | 0.707517 | 10 |
| 5 | 1820.0 | 3528.0 | 1708.0 | 0.076221 | 0.0 | 0.000010 | 0.0 | 922.273682 | 0.539973 | 0.076221 | ... | 0.0 | 1457.363647 | 0.853257 | 0.120443 | 0.0 | 0.000201 | 0.0 | 1457.363647 | 0.853257 | 11 |
| 6 | 2014.0 | 4259.0 | 2245.0 | 0.080958 | 0.0 | 0.008331 | 0.0 | 979.588989 | 0.436343 | 0.080958 | ... | 0.0 | 1587.135742 | 0.706965 | 0.131168 | 0.0 | 0.135986 | 0.0 | 1587.135742 | 0.706965 | 13 |
7 rows × 58 columns