Sharded Cellpose Segmentation#
[ ]:
import os
import os
import numpy as np
from scportrait.pipeline.featurization import CellFeaturizer
from scportrait.pipeline.extraction import HDF5CellExtraction
from scportrait.pipeline.project import Project
from scportrait.pipeline.segmentation.workflows import ShardedCytosolSegmentationCellpose
from scportrait.pipeline.selection import LMDSelection
import scportrait
[2]:
project_location = "project_sharded"
config_path = scportrait.data.get_config_file(config_id = "dataset_1_config")
project = Project(
os.path.abspath(project_location),
config_path=config_path,
overwrite=True,
debug=True,
segmentation_f=ShardedCytosolSegmentationCellpose,
extraction_f=HDF5CellExtraction,
featurization_f=CellFeaturizer,
selection_f=LMDSelection,
)
Updating project config file.
[10/04/2025 18:39:08] Loading config from /Users/sophia/Documents/GitHub/scPortrait/examples/notebooks/example_projects/example_1/project_sharded/config.yml
[10/04/2025 18:39:08] Initialized temporary directory at /var/folders/35/p4c58_4n3bb0bxnzgns1t7kh0000gn/T/./ShardedCytosolSegmentationCellpose_tq5kean9 for ShardedCytosolSegmentationCellpose
[10/04/2025 18:39:08] Compression algorithm for extracted single-cell images: lzf
[3]:
dataset_1_path = scportrait.data.dataset_1()
# these example images are downloaded from the human protein atlas (www.proteinatlas.org)
images = [f"{dataset_1_path}/Ch1.tif", f"{dataset_1_path}/Ch2.tif", f"{dataset_1_path }/Ch3.tif"]
project.load_input_from_tif_files(images, channel_names = ["Hoechst", "Alexa488", "mCherry"])
[10/04/2025 18:39:21] Output location /Users/sophia/Documents/GitHub/scPortrait/examples/notebooks/example_projects/example_1/project_sharded/scportrait.sdata already exists. Overwriting.
INFO The Zarr backing store has been changed from None the new file path:
/Users/sophia/Documents/GitHub/scPortrait/examples/notebooks/example_projects/example_1/project_sharded/sc
portrait.sdata
[10/04/2025 18:39:21] Initialized temporary directory at /Users/sophia/Documents/GitHub/scPortrait/examples/notebooks/example_projects/example_1/Project_ol63yoc2 for Project
[10/04/2025 18:39:22] Image input_image written to sdata object.
[10/04/2025 18:39:22] Cleaned up temporary directory at <TemporaryDirectory '/Users/sophia/Documents/GitHub/scPortrait/examples/notebooks/example_projects/example_1/Project_ol63yoc2'>
[4]:
project.plot_input_image()
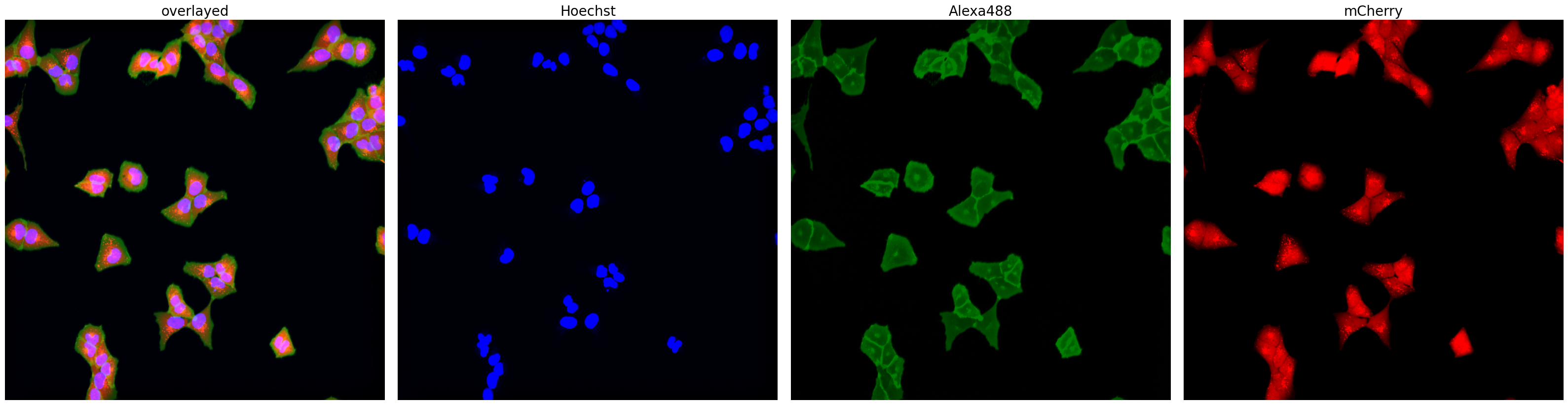
[5]:
project.segment()
[10/04/2025 18:39:24] Mapped input image to memory-mapped array.
[10/04/2025 18:39:24] Created new shard directory /Users/sophia/Documents/GitHub/scPortrait/examples/notebooks/example_projects/example_1/project_sharded/segmentation/tiles
[10/04/2025 18:39:24] sharding plan already found in directory /Users/sophia/Documents/GitHub/scPortrait/examples/notebooks/example_projects/example_1/project_sharded/segmentation/sharding_plan.csv.
[10/04/2025 18:39:24] Overwriting existing sharding plan.
[10/04/2025 18:39:24] target size 2000000 is smaller than input image 9229443. Sharding will be used.
[10/04/2025 18:39:24] input image 3039 px by 3037 px
[10/04/2025 18:39:24] target_shard_size: 2000000
[10/04/2025 18:39:24] sharding plan:
[10/04/2025 18:39:24] 2 rows by 2 columns
[10/04/2025 18:39:24] 1519 px by 1518 px
[10/04/2025 18:39:24] Saving Sharding plan to file: /Users/sophia/Documents/GitHub/scPortrait/examples/notebooks/example_projects/example_1/project_sharded/segmentation/sharding_plan.csv
[10/04/2025 18:39:24] Initialized temporary directory at /var/folders/35/p4c58_4n3bb0bxnzgns1t7kh0000gn/T/./CytosolSegmentationCellpose_8s21a409 for CytosolSegmentationCellpose
[10/04/2025 18:39:24] Initialized temporary directory at /var/folders/35/p4c58_4n3bb0bxnzgns1t7kh0000gn/T/./CytosolSegmentationCellpose_dcusqple for CytosolSegmentationCellpose
[10/04/2025 18:39:24] Initialized temporary directory at /var/folders/35/p4c58_4n3bb0bxnzgns1t7kh0000gn/T/./CytosolSegmentationCellpose_79uimt4e for CytosolSegmentationCellpose
[10/04/2025 18:39:24] Initialized temporary directory at /var/folders/35/p4c58_4n3bb0bxnzgns1t7kh0000gn/T/./CytosolSegmentationCellpose_1okhswgp for CytosolSegmentationCellpose
[10/04/2025 18:39:24] sharding plan with 4 elements generated, sharding with 2 threads begins
[10/04/2025 18:39:24] GPU Status for segmentation is True with 1 GPUs found. Segmentation will be performed on the device mps with 2 processes per device in parallel.
[10/04/2025 18:39:30] Beginning Segmentation of Shard with the slicing (slice(0, 1619, None), slice(0, 1618, None))
[10/04/2025 18:39:30] Beginning Segmentation of Shard with the slicing (slice(0, 1619, None), slice(1418, 3037, None))
[10/04/2025 18:39:30] Time taken to load input image: 0.0055277499777730554
[10/04/2025 18:39:30] Time taken to load input image: 0.005530958995223045
[10/04/2025 18:39:30] GPU Status for segmentation is True and will segment using the following device mps.
[10/04/2025 18:39:30] GPU Status for segmentation is True and will segment using the following device mps.
[10/04/2025 18:39:30] Segmenting nucleus using the following model: nuclei[10/04/2025 18:39:30] Segmenting nucleus using the following model: nuclei
[10/04/2025 18:39:35] Segmenting cytosol using the following model: cyto2
[10/04/2025 18:39:35] Segmenting cytosol using the following model: cyto2
[10/04/2025 18:39:41] Performing filtering to match Cytosol and Nucleus IDs.
[10/04/2025 18:39:41] Performing filtering to match Cytosol and Nucleus IDs.
[10/04/2025 18:39:42] Removed 46 nuclei and 24 cytosols due to filtering.
[10/04/2025 18:39:42] After filtering, 109 matching nuclei and cytosol masks remain.
[10/04/2025 18:39:42] Removed 40 nuclei and 16 cytosols due to filtering.
[10/04/2025 18:39:42] After filtering, 152 matching nuclei and cytosol masks remain.
[10/04/2025 18:39:44] Total time to perform nucleus and cytosol mask matching filtering: 3.12 seconds
[10/04/2025 18:39:44] Filtering status for this segmentation is set to True.
[10/04/2025 18:39:44] Filtering has been performed during segmentation. Nucleus and Cytosol IDs match. No additional steps are required.
[10/04/2025 18:39:44] Saved cell_id classes to file /Users/sophia/Documents/GitHub/scPortrait/examples/notebooks/example_projects/example_1/project_sharded/segmentation/tiles/1/classes.csv.
[10/04/2025 18:39:44] === Finished segmentation of shard ===
[10/04/2025 18:39:44] Cleaned up temporary directory at <TemporaryDirectory '/var/folders/35/p4c58_4n3bb0bxnzgns1t7kh0000gn/T/./CytosolSegmentationCellpose_dcusqple'>
[10/04/2025 18:39:44] Total time to perform nucleus and cytosol mask matching filtering: 3.38 seconds
[10/04/2025 18:39:44] Segmentation of Shard with the slicing (slice(0, 1619, None), slice(1418, 3037, None)) finished
[10/04/2025 18:39:44] Beginning Segmentation of Shard with the slicing (slice(1419, 3039, None), slice(0, 1618, None))
[10/04/2025 18:39:44] Time taken to load input image: 0.0061804589931853116
[10/04/2025 18:39:44] Filtering status for this segmentation is set to True.
[10/04/2025 18:39:44] Filtering has been performed during segmentation. Nucleus and Cytosol IDs match. No additional steps are required.
[10/04/2025 18:39:44] Saved cell_id classes to file /Users/sophia/Documents/GitHub/scPortrait/examples/notebooks/example_projects/example_1/project_sharded/segmentation/tiles/0/classes.csv.
[10/04/2025 18:39:44] === Finished segmentation of shard ===
[10/04/2025 18:39:44] GPU Status for segmentation is True and will segment using the following device mps.
[10/04/2025 18:39:45] Cleaned up temporary directory at <TemporaryDirectory '/var/folders/35/p4c58_4n3bb0bxnzgns1t7kh0000gn/T/./CytosolSegmentationCellpose_8s21a409'>
[10/04/2025 18:39:45] Segmentation of Shard with the slicing (slice(0, 1619, None), slice(0, 1618, None)) finished
[10/04/2025 18:39:45] Beginning Segmentation of Shard with the slicing (slice(1419, 3039, None), slice(1418, 3037, None))
[10/04/2025 18:39:45] Time taken to load input image: 0.005262999999104068
[10/04/2025 18:39:45] GPU Status for segmentation is True and will segment using the following device mps.
[10/04/2025 18:39:45] Segmenting nucleus using the following model: nuclei
[10/04/2025 18:39:45] Segmenting nucleus using the following model: nuclei
[10/04/2025 18:39:47] Segmenting cytosol using the following model: cyto2
[10/04/2025 18:39:47] Segmenting cytosol using the following model: cyto2
[10/04/2025 18:39:52] Performing filtering to match Cytosol and Nucleus IDs.
[10/04/2025 18:39:52] Performing filtering to match Cytosol and Nucleus IDs.
[10/04/2025 18:39:52] Removed 33 nuclei and 13 cytosols due to filtering.
[10/04/2025 18:39:52] After filtering, 70 matching nuclei and cytosol masks remain.
[10/04/2025 18:39:53] Removed 59 nuclei and 25 cytosols due to filtering.
[10/04/2025 18:39:53] After filtering, 88 matching nuclei and cytosol masks remain.
[10/04/2025 18:39:54] Total time to perform nucleus and cytosol mask matching filtering: 2.63 seconds
[10/04/2025 18:39:54] Filtering status for this segmentation is set to True.
[10/04/2025 18:39:54] Filtering has been performed during segmentation. Nucleus and Cytosol IDs match. No additional steps are required.
[10/04/2025 18:39:54] Saved cell_id classes to file /Users/sophia/Documents/GitHub/scPortrait/examples/notebooks/example_projects/example_1/project_sharded/segmentation/tiles/3/classes.csv.
[10/04/2025 18:39:54] === Finished segmentation of shard ===
[10/04/2025 18:39:55] Cleaned up temporary directory at <TemporaryDirectory '/var/folders/35/p4c58_4n3bb0bxnzgns1t7kh0000gn/T/./CytosolSegmentationCellpose_1okhswgp'>
[10/04/2025 18:39:55] Segmentation of Shard with the slicing (slice(1419, 3039, None), slice(1418, 3037, None)) finished
[10/04/2025 18:39:55] Total time to perform nucleus and cytosol mask matching filtering: 2.94 seconds
[10/04/2025 18:39:55] Filtering status for this segmentation is set to True.
[10/04/2025 18:39:55] Filtering has been performed during segmentation. Nucleus and Cytosol IDs match. No additional steps are required.
[10/04/2025 18:39:55] Saved cell_id classes to file /Users/sophia/Documents/GitHub/scPortrait/examples/notebooks/example_projects/example_1/project_sharded/segmentation/tiles/2/classes.csv.
[10/04/2025 18:39:55] === Finished segmentation of shard ===
[10/04/2025 18:39:55] Cleaned up temporary directory at <TemporaryDirectory '/var/folders/35/p4c58_4n3bb0bxnzgns1t7kh0000gn/T/./CytosolSegmentationCellpose_79uimt4e'>
[10/04/2025 18:39:55] Segmentation of Shard with the slicing (slice(1419, 3039, None), slice(0, 1618, None)) finished
[10/04/2025 18:39:56] Finished parallel segmentation
[10/04/2025 18:39:56] resolve sharding plan
[10/04/2025 18:39:56] Cleared temporary directory containing input image used for sharding.
[10/04/2025 18:39:56] Stitching tile 0
[10/04/2025 18:39:57] Time taken to cleanup overlapping shard regions for shard 0: 0.28040599822998047s
[10/04/2025 18:39:57] Number of classes contained in shard after processing: 152
[10/04/2025 18:39:57] Number of Ids in filtered_classes after adding shard 0: 152
[10/04/2025 18:39:57] Finished stitching tile 0 in 1.044527292251587 seconds.
[10/04/2025 18:39:57] Number of filtered classes in Dataset: 152
[10/04/2025 18:39:57] Filtering status for this segmentation is set to True.
[10/04/2025 18:39:57] Filtering has been performed during segmentation. Nucleus and Cytosol IDs match. No additional steps are required.
[10/04/2025 18:39:57] Saved cell_id classes to file /Users/sophia/Documents/GitHub/scPortrait/examples/notebooks/example_projects/example_1/project_sharded/segmentation/classes.csv.
[10/04/2025 18:39:57] Stitching tile 1
[10/04/2025 18:39:57] Time taken to cleanup overlapping shard regions for shard 1: 0.0597681999206543s
[10/04/2025 18:39:57] Number of classes contained in shard after processing: 112
[10/04/2025 18:39:57] Number of Ids in filtered_classes after adding shard 1: 241
[10/04/2025 18:39:57] Finished stitching tile 1 in 0.12884902954101562 seconds.
[10/04/2025 18:39:57] Number of filtered classes in Dataset: 241
[10/04/2025 18:39:57] Filtering status for this segmentation is set to True.
[10/04/2025 18:39:57] Filtering has been performed during segmentation. Nucleus and Cytosol IDs match. No additional steps are required.
[10/04/2025 18:39:57] Saved cell_id classes to file /Users/sophia/Documents/GitHub/scPortrait/examples/notebooks/example_projects/example_1/project_sharded/segmentation/classes.csv.
[10/04/2025 18:39:57] Stitching tile 2
[10/04/2025 18:39:58] Time taken to cleanup overlapping shard regions for shard 2: 0.07414388656616211s
[10/04/2025 18:39:58] Number of classes contained in shard after processing: 99
[10/04/2025 18:39:58] Number of Ids in filtered_classes after adding shard 2: 324
[10/04/2025 18:39:58] Finished stitching tile 2 in 0.14506816864013672 seconds.
[10/04/2025 18:39:58] Number of filtered classes in Dataset: 324
[10/04/2025 18:39:58] Filtering status for this segmentation is set to True.
[10/04/2025 18:39:58] Filtering has been performed during segmentation. Nucleus and Cytosol IDs match. No additional steps are required.
[10/04/2025 18:39:58] Saved cell_id classes to file /Users/sophia/Documents/GitHub/scPortrait/examples/notebooks/example_projects/example_1/project_sharded/segmentation/classes.csv.
[10/04/2025 18:39:58] Stitching tile 3
[10/04/2025 18:39:58] Time taken to cleanup overlapping shard regions for shard 3: 0.05631685256958008s
[10/04/2025 18:39:58] Number of classes contained in shard after processing: 74
[10/04/2025 18:39:58] Number of Ids in filtered_classes after adding shard 3: 379
[10/04/2025 18:39:58] Finished stitching tile 3 in 0.12271714210510254 seconds.
[10/04/2025 18:39:58] Number of filtered classes in Dataset: 379
[10/04/2025 18:39:58] Filtering status for this segmentation is set to True.
[10/04/2025 18:39:58] Filtering has been performed during segmentation. Nucleus and Cytosol IDs match. No additional steps are required.
[10/04/2025 18:39:58] Saved cell_id classes to file /Users/sophia/Documents/GitHub/scPortrait/examples/notebooks/example_projects/example_1/project_sharded/segmentation/classes.csv.
[10/04/2025 18:39:58] resolved sharding plan.
[10/04/2025 18:39:58] Segmentation seg_all_nucleus written to sdata object.
[10/04/2025 18:40:00] Points centers_seg_all_nucleus written to sdata object.
[10/04/2025 18:40:00] Segmentation seg_all_cytosol written to sdata object.
[10/04/2025 18:40:00] Points centers_seg_all_cytosol written to sdata object.
[10/04/2025 18:40:01] Cleaned up temporary directory at <TemporaryDirectory '/var/folders/35/p4c58_4n3bb0bxnzgns1t7kh0000gn/T/./ShardedCytosolSegmentationCellpose_tq5kean9'>
[10/04/2025 18:40:01] finished saving segmentation results to sdata object for sharded segmentation.
[10/04/2025 18:40:01] Deleting intermediate tile results to free up storage space
[10/04/2025 18:40:01] Total time taken for sharded segmentation: 36.54483179200906 seconds
[10/04/2025 18:40:01] === finished sharded segmentation ===
[6]:
project.plot_segmentation_masks()
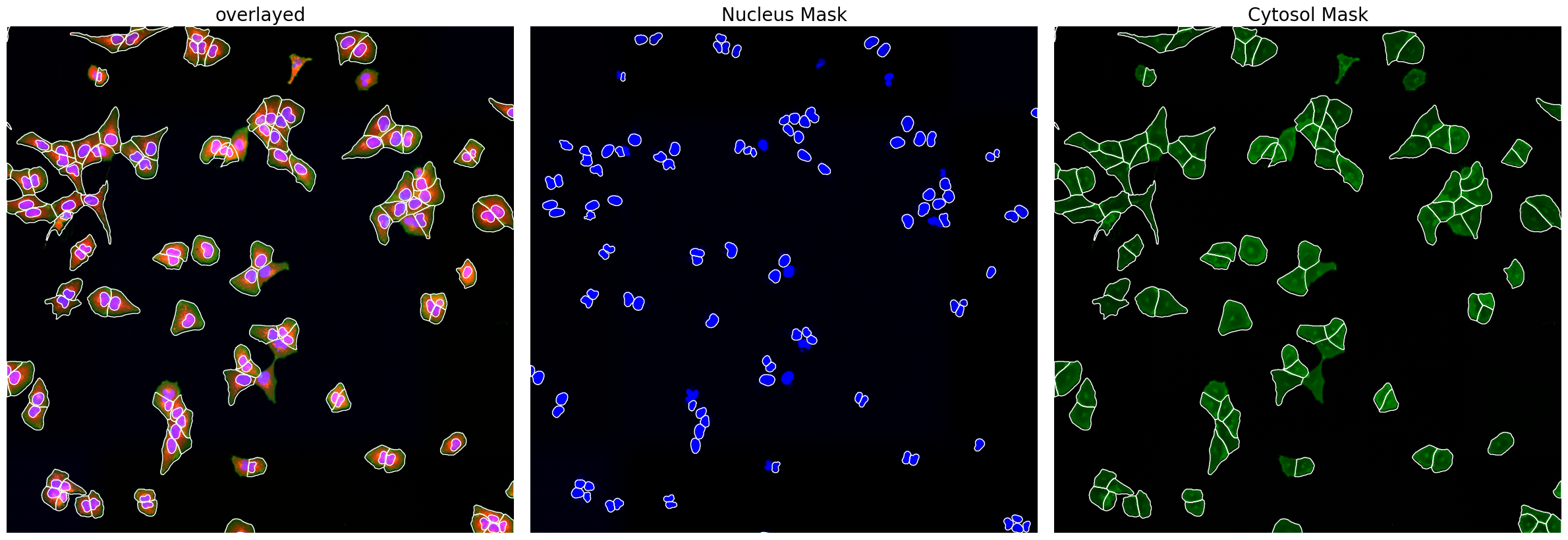
[7]:
project.extract()
[10/04/2025 18:40:08] Initialized temporary directory at /var/folders/35/p4c58_4n3bb0bxnzgns1t7kh0000gn/T/./HDF5CellExtraction_fe448s0x for HDF5CellExtraction
[10/04/2025 18:40:08] Created new directory for extraction results: /Users/sophia/Documents/GitHub/scPortrait/examples/notebooks/example_projects/example_1/project_sharded/extraction/data
[10/04/2025 18:40:08] Setup output folder at /Users/sophia/Documents/GitHub/scPortrait/examples/notebooks/example_projects/example_1/project_sharded/extraction/data
[10/04/2025 18:40:08] Found 2 segmentation masks for the given key in the sdata object. Will be extracting single-cell images based on these masks: ['seg_all_nucleus', 'seg_all_cytosol']
[10/04/2025 18:40:08] Using seg_all_nucleus as the main segmentation mask to determine cell centers.
[10/04/2025 18:40:08] A total of 9 cells were too close to the image border to be extracted. Their cell_ids were saved to file /Users/sophia/Documents/GitHub/scPortrait/examples/notebooks/example_projects/example_1/project_sharded/extraction/removed_classes.csv.
[10/04/2025 18:40:08] Container for single-cell data created.
[10/04/2025 18:40:08] Extraction Details:
[10/04/2025 18:40:08] --------------------------------
[10/04/2025 18:40:08] Number of input image channels: 3
[10/04/2025 18:40:08] Number of segmentation masks used during extraction: 2
[10/04/2025 18:40:08] Number of generated output images per cell: 5
[10/04/2025 18:40:08] Number of unique cells to extract: 370
[10/04/2025 18:40:08] Extracted Image Dimensions: 128 x 128
[10/04/2025 18:40:08] Normalization of extracted images: True
[10/04/2025 18:40:08] Percentile normalization range for single-cell images: [0.01, 0.99]
[10/04/2025 18:40:08] Starting single-cell image extraction of 370 cells...
[10/04/2025 18:40:08] Loading input images to memory mapped arrays...
[10/04/2025 18:40:09] Finished transferring data to memory mapped arrays. Time taken: 0.46 seconds.
[10/04/2025 18:40:09] Using batch size of 100 for multiprocessing.
[10/04/2025 18:40:09] Running in multiprocessing mode with 4 threads.
[10/04/2025 18:40:09] Finished extraction in 0.50 seconds (732.72 cells / second)
[10/04/2025 18:40:10] Benchmarking times saved to file.
[10/04/2025 18:40:10] Cleaned up temporary directory at <TemporaryDirectory '/var/folders/35/p4c58_4n3bb0bxnzgns1t7kh0000gn/T/./HDF5CellExtraction_fe448s0x'>
[8]:
project.plot_single_cell_images()
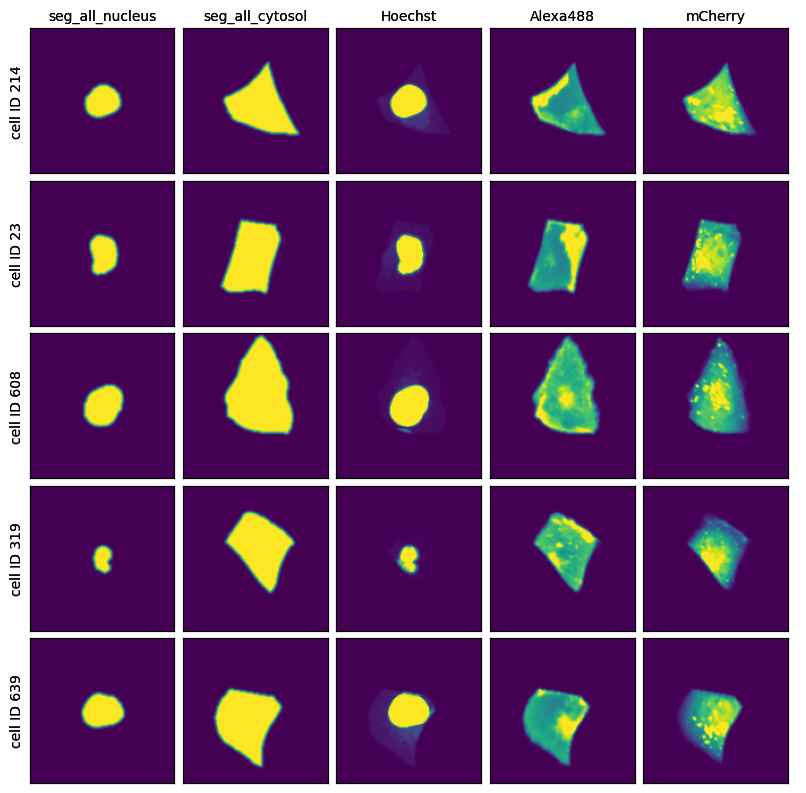
[9]:
project.featurize(overwrite = True)
Using extraction directory: /Users/sophia/Documents/GitHub/scPortrait/examples/notebooks/example_projects/example_1/project_sharded/extraction/data/single_cells.h5sc
[10/04/2025 18:40:12] Initialized temporary directory at /var/folders/35/p4c58_4n3bb0bxnzgns1t7kh0000gn/T/./CellFeaturizer_fz3haljn for CellFeaturizer
[10/04/2025 18:40:12] Started CellFeaturization of all available channels.
[10/04/2025 18:40:12] Created new directory for featurization results: /Users/sophia/Documents/GitHub/scPortrait/examples/notebooks/example_projects/example_1/project_sharded/featurization/complete_CellFeaturizer
[10/04/2025 18:40:12] CPU specified in config file but MPS available on system. Consider changing the device for the next run.
[10/04/2025 18:40:12] Initialized temporary directory at /var/folders/35/p4c58_4n3bb0bxnzgns1t7kh0000gn/T/./CellFeaturizer_j3rqkviq for CellFeaturizer
[10/04/2025 18:40:12] Reading data from path: /Users/sophia/Documents/GitHub/scPortrait/examples/notebooks/example_projects/example_1/project_sharded/extraction/data/single_cells.h5sc
[10/04/2025 18:40:12] Processing dataset with 370 cells
[10/04/2025 18:40:12] Dataloader generated with a batchsize of 900 and 10 workers. Dataloader contains 1 entries.
[10/04/2025 18:40:12] Started processing of 1 batches.
[10/04/2025 18:40:14] finished processing.
[10/04/2025 18:40:14] Results saved to file: /Users/sophia/Documents/GitHub/scPortrait/examples/notebooks/example_projects/example_1/project_sharded/featurization/complete_CellFeaturizer/calculated_image_features.csv
[10/04/2025 18:40:14] Table CellFeaturizer_nucleus written to sdata object.
[10/04/2025 18:40:14] Table CellFeaturizer_cytosol written to sdata object.
[10/04/2025 18:40:14] GPU memory before performing cleanup: None
[10/04/2025 18:40:14] GPU memory after performing cleanup: None
[10/04/2025 18:40:15] Cleaned up temporary directory at <TemporaryDirectory '/var/folders/35/p4c58_4n3bb0bxnzgns1t7kh0000gn/T/./CellFeaturizer_j3rqkviq'>
[10]:
# load classification results
results = project.sdata['CellFeaturizer_cytosol'].to_df().merge(project.sdata['CellFeaturizer_cytosol'].obs, left_index=True, right_index=True).drop(columns = "region")
results
[10]:
| nucleus_area | cytosol_area | cytosol_only_area | Hoechst_mean_nucleus | Hoechst_median_nucleus | Hoechst_quant75_nucleus | Hoechst_quant25_nucleus | Hoechst_summed_intensity_nucleus | Hoechst_summed_intensity_area_normalized_nucleus | Hoechst_mean_cytosol | ... | mCherry_quant25_cytosol | mCherry_summed_intensity_cytosol | mCherry_summed_intensity_area_normalized_cytosol | mCherry_mean_cytosol_only | mCherry_median_cytosol_only | mCherry_quant75_cytosol_only | mCherry_quant25_cytosol_only | mCherry_summed_intensity_cytosol_only | mCherry_summed_intensity_area_normalized_cytosol_only | scportrait_cell_id | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 1275.0 | 4273.0 | 2998.0 | 0.040320 | 0.0 | 1.192093e-07 | 0.0 | 660.602295 | 0.220348 | 0.040320 | ... | 0.0 | 1916.359619 | 0.639213 | 0.116965 | 0.0 | 2.980232e-07 | 0.0 | 1916.359619 | 0.639213 | 14 |
| 1 | 1381.0 | 3870.0 | 2489.0 | 0.055529 | 0.0 | 0.000000e+00 | 0.0 | 909.779480 | 0.365520 | 0.055529 | ... | 0.0 | 1684.037476 | 0.676592 | 0.102785 | 0.0 | 0.000000e+00 | 0.0 | 1684.037476 | 0.676592 | 15 |
| 2 | 1395.0 | 4980.0 | 3585.0 | 0.057506 | 0.0 | 1.778841e-03 | 0.0 | 942.177368 | 0.262811 | 0.057506 | ... | 0.0 | 2160.976562 | 0.602783 | 0.131896 | 0.0 | 3.061295e-03 | 0.0 | 2160.976562 | 0.602783 | 16 |
| 3 | 1486.0 | 4273.0 | 2799.0 | 0.059866 | 0.0 | 1.072884e-06 | 0.0 | 980.838196 | 0.350424 | 0.059866 | ... | 0.0 | 1751.940918 | 0.625917 | 0.106930 | 0.0 | 6.556511e-07 | 0.0 | 1751.940918 | 0.625917 | 17 |
| 4 | 1706.0 | 6824.0 | 5118.0 | 0.073903 | 0.0 | 2.632141e-02 | 0.0 | 1210.828613 | 0.236582 | 0.073903 | ... | 0.0 | 2234.593750 | 0.436615 | 0.136389 | 0.0 | 1.261292e-01 | 0.0 | 2234.593750 | 0.436615 | 18 |
| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
| 365 | 1054.0 | 2744.0 | 1696.0 | 0.033506 | 0.0 | 0.000000e+00 | 0.0 | 548.970093 | 0.323685 | 0.033506 | ... | 0.0 | 1313.792725 | 0.774642 | 0.080188 | 0.0 | 0.000000e+00 | 0.0 | 1313.792725 | 0.774642 | 1339 |
| 366 | 1468.0 | 5968.0 | 4500.0 | 0.055512 | 0.0 | 2.802849e-03 | 0.0 | 909.506470 | 0.202113 | 0.055512 | ... | 0.0 | 2116.351562 | 0.470300 | 0.129172 | 0.0 | 5.178070e-02 | 0.0 | 2116.351562 | 0.470300 | 1340 |
| 367 | 1124.0 | 4149.0 | 3025.0 | 0.034669 | 0.0 | 0.000000e+00 | 0.0 | 568.015991 | 0.187774 | 0.034669 | ... | 0.0 | 1566.534424 | 0.517863 | 0.095614 | 0.0 | 0.000000e+00 | 0.0 | 1566.534424 | 0.517863 | 1341 |
| 368 | 1498.0 | 6155.0 | 4657.0 | 0.055164 | 0.0 | 2.731323e-03 | 0.0 | 903.804321 | 0.194074 | 0.055164 | ... | 0.0 | 2243.846680 | 0.481822 | 0.136954 | 0.0 | 6.857300e-02 | 0.0 | 2243.846680 | 0.481822 | 1342 |
| 369 | 1196.0 | 5577.0 | 4381.0 | 0.038588 | 0.0 | 1.783848e-03 | 0.0 | 632.223816 | 0.144310 | 0.038588 | ... | 0.0 | 2038.879517 | 0.465391 | 0.124443 | 0.0 | 1.167488e-02 | 0.0 | 2038.879517 | 0.465391 | 1343 |
370 rows × 58 columns
[12]:
#test selection workflow
selected_cells_large = results[results.cytosol_area > 4500]["scportrait_cell_id"].tolist()
selected_cells_small = results[results.cytosol_area < 3000]["scportrait_cell_id"].tolist()
cells_to_select = [
{"name": "large_cells", "classes": selected_cells_large, "well": "A1"},
{"name": "small_cells", "classes": selected_cells_small, "well": "B1"},
]
marker_0 = (0, 0)
marker_1 = (2000, 0)
marker_2 = (0, 2000)
calibration_marker = np.array([marker_0, marker_1, marker_2])
[13]:
project.select(cells_to_select, calibration_marker)
[10/04/2025 18:41:12] Initialized temporary directory at /var/folders/35/p4c58_4n3bb0bxnzgns1t7kh0000gn/T/./LMDSelection_rllzvk72 for LMDSelection
[10/04/2025 18:41:12] Selection process started.
[10/04/2025 18:41:21] Temporary directory not found, skipping cleanup
[10/04/2025 18:41:21] Temporary directory not found, skipping cleanup
[10/04/2025 18:41:21] Temporary directory not found, skipping cleanup
[10/04/2025 18:41:21] Coordinate lookup index calculation took 8.985896790982224 seconds.
No configuration for shape_erosion found, parameter will be set to 0
No configuration for binary_smoothing found, parameter will be set to 3
No configuration for convolution_smoothing found, parameter will be set to 15
No configuration for rdp_epsilon found, parameter will be set to 0.1
No configuration for xml_decimal_transform found, parameter will be set to 100
No configuration for distance_heuristic found, parameter will be set to 300
No configuration for join_intersecting found, parameter will be set to True
Path optimizer used for XML generation: hilbert
cell set 0 passed sanity check
cell set 1 passed sanity check
Loading coordinates from external source
Processing cell sets in parallel
Convert label format into coordinate format
Conversion finished, performing sanity check.
Intersecting Shapes will be merged into a single shape.
Convert label format into coordinate format
Conversion finished, performing sanity check.
Intersecting Shapes will be merged into a single shape.
dilating shapes: 100%|██████████| 43/43 [00:04<00:00, 9.21it/s]
dilating shapes: 100%|██████████| 169/169 [00:05<00:00, 32.43it/s]
0 shapes that were intersecting were found and merged.
creating shapes: 0%| | 0/31 [00:00<?, ?it/s]
0 shapes that were intersecting were found and merged.
creating shapes: 100%|██████████| 31/31 [00:04<00:00, 6.97it/s]
creating shapes: 100%|██████████| 106/106 [00:05<00:00, 19.56it/s]
calculating polygons: 100%|██████████| 31/31 [00:06<00:00, 4.99it/s]
Current path length: 21,152.45 units
calculating polygons: 2%|▏ | 2/106 [00:04<02:57, 1.70s/it]
Optimized path length: 14,705.91 units
Optimization factor: 1.4x
calculating polygons: 5%|▍ | 5/106 [00:04<00:57, 1.77it/s]
Plotting shapes in debug mode is not supported in multi-threading mode.
Saving plots to disk instead.
calculating polygons: 100%|██████████| 106/106 [00:05<00:00, 17.78it/s]
Current path length: 59,053.81 units
Optimized path length: 28,792.09 units
Optimization factor: 2.1x
Plotting shapes in debug mode is not supported in multi-threading mode.
Saving plots to disk instead.
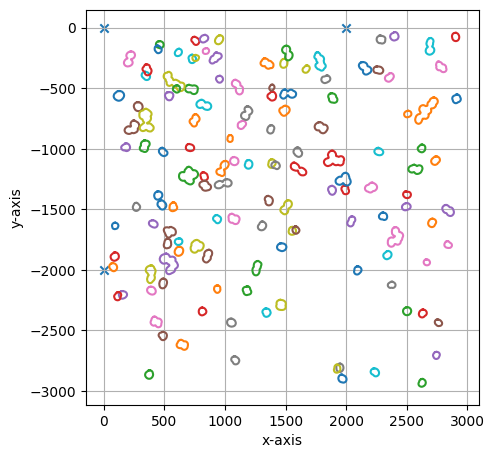
===== Collection Stats =====
Number of shapes: 137
Number of vertices: 10,753
============================
Mean vertices: 78
Min vertices: 52
5% percentile vertices: 57
Median vertices: 67
95% percentile vertices: 141
Max vertices: 215
[0 0]
[ 0 -200000]
[200000 0]
[10/04/2025 18:41:43] Saved output at /Users/sophia/Documents/GitHub/scPortrait/examples/notebooks/example_projects/example_1/project_sharded/selection/large_cells_small_cells.xml
[10/04/2025 18:41:43] Cleaned up temporary directory at <TemporaryDirectory '/var/folders/35/p4c58_4n3bb0bxnzgns1t7kh0000gn/T/./LMDSelection_rllzvk72'>
[ ]: